Which tools can I use to share my data on GBIF?
As you probably already know, GBIF.org doesn’t host any data. The system relies on each data provider making their data available online in a GBIF-supported format. It also relies on organization letting GBIF know where to find these data (in other words registering the data). But how to do just that?
The good news is that there are several GBIF-compatible systems. They will export or make available the data for you in the correct format and several provide means to register them as datasets on GBIF.
In this blogpost, I will try to give a very brief overview of these tools and how they work with GBIF.
If you are new to GBIF you might want to read our Quick guide to publishing data through GBIF.org first. In addition to that, I would also suggest that you take a look at our documentation concerning data hosting.
Types of entities on GBIF: datasets, organizations, nodes, installations
Before I start listing the various types of GBIF-compatible portals and systems, I need to explain some of the GBIF vocabulary.
We have four kind of entities on GBIF (or four kinds of web pages if you prefer):
- Datasets (for example: the Prairie Fen Biodiversity Project).
- These datasets are published by an organization - also called publisher (for example: the Central Michigan University Herbarium).
- Each organization is endorsed by a node (for example: the U.S. Geological Survey) or a committee (for organizations in non-participating countries). Endorsement not only ensures suitable data is shared, but often helps make connections between people in the network who can assist each other.
- These datasets are also hosted by an installation (for example: IPT - Hosted by iDigBio). An installation is attached to an organization. The type of an installation will depend on the type of publishing tool/portal/system you choose.
I tried to summarize all of this in the schema below:
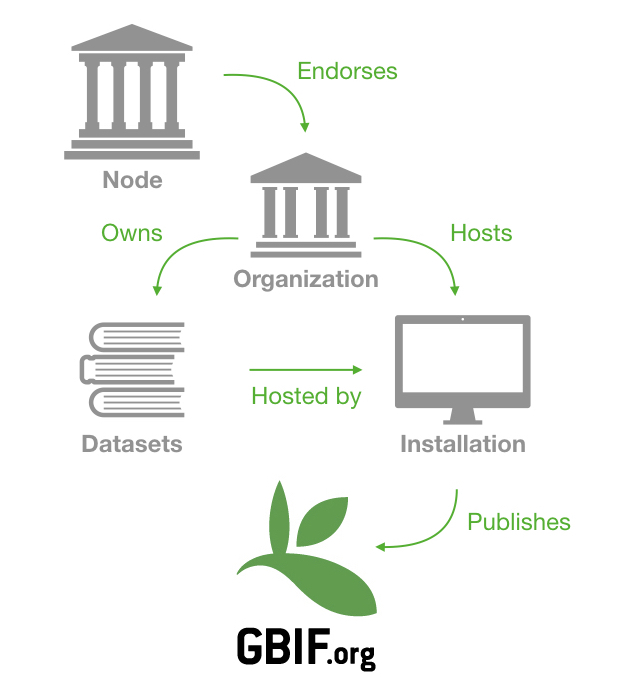
But of course there are some variants to schema. The organization owning the data doesn’t have to be the same as the organization hosting the data. For example, the Prairie Fen Biodiversity Project is published by the Central Michigan University Herbarium but hosted by an installation attached to the Arizona State University Biodiversity Knowledge Integration Center.
In that case, we will differentiate Publishing Organization and Hosting organization (see the schema below). Both organizations can also be endorsed by different nodes.
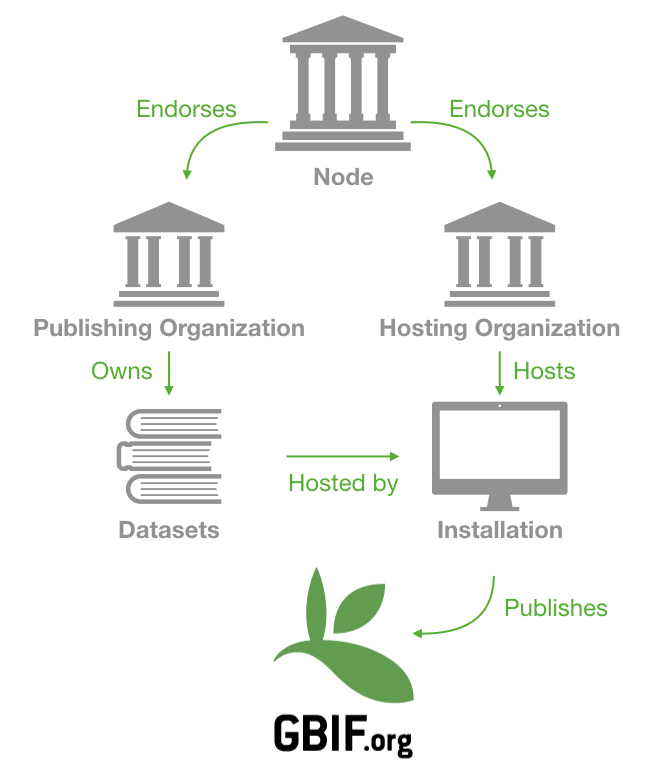
All of these entities are registered on GBIF. Amongst other things, this structure aims to represent everyone’s contribution to the data and allow us to aggregate statistics at different levels.
What if you are not affiliated with any organization? You can take part in one of the citizen science initiatives which contribute to GBIF such as iNaturalist.
IPT: The Integrated Publishing Toolkit
The IPT is the tool developed and maintained by the GBIF secretariat. It is the most common type of installation connected to GBIF and provides the repository network for the marine community OBIS. IPTs can generate Darwin Core Archives for each dataset and register them on GBIF.
The IPT was designed to share data on GBIF, it is not a data management system. However, some actual data management systems can export data to IPTs. This is the case for Specify, a software to manage specimen data for biological research collections.
Setting up hosting organization
When setting up an IPT, you will be asked to associate it with a (hosting) organization.
Setting up publishing organizations
- When an organization is endorsed, helpdesk@gbif.org will send to the organization technical contact(s) an IPT token associated with this organization (also called “organization password”).
- The token will be needed to add the organization to an IPT. Only a IPT administrator can add an organization to an IPT. There is no limit on the number of organization registered in an IPT.
- IPT users can then select the relevant publishing organization when creating a publishing a new dataset.
For more information, check out the IPT documentation.
API-based systems
What I call “API-based systems” are tools/portals that rely on the GBIF registry API to share data on GBIF. Most of these tools are data-management systems. They usually have many features and being GBIF-compatible is only one of them. Here are some of these tools (in no particular order):
- The Living Atlases: Open source IT infrastructure for the aggregation and delivery of biodiversity data.
- Symbiota: Open source content management system for curating specimen- and observation-based biodiversity data.
- PlutoF: Data management and Publishing Platform
- Earthcape: Biodiversity database platform.
- Others: some organizations such as the Naturalis Biodiversity Center also made their own system GBIF-compatible by using our API.
Setting up hosting and publishing organizations
These systems generate and host Darwin Core Archives and register their endpoints on GBIF using the API. Because they use the registry API, all of them are associated with an API user account (= API username). This API username can be given permissions to register and update datasets on behalf of an organization or a node.
If a username is given permissions to register and update datasets for a node, all the datasets from the organizations endorsed by this Node are concerned (this is the case for the NBN Atlas for example). However, in most cases, the permissions are set for one organization at the time.
The GBIF Secretariat handle these permissions. This means that organizations have to send an email to heldpesk@gbif.org to request that a specific API username is given the permission to register and update their datasets.
I tried to summarize all of this below:
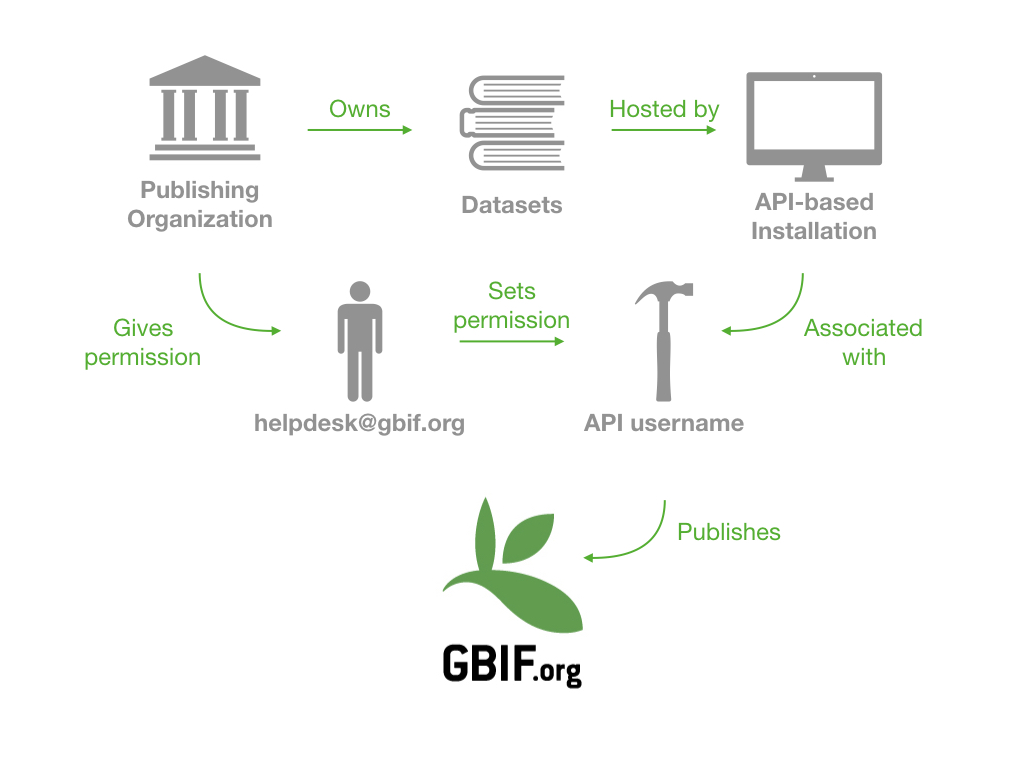
Other systems
Some systems expose databases online through open web protocols, which require setting up some specific wrappers. As for the API-based systems, being GBIF-compatible is only one of their features. Here are some of these tools:
- BioCASe (BioCASe and BioCASe Provider Software are more than that, see the next paragraph for some explanations)
- DiGIR (still GBIF-compatible but has no active support)
- TAPIR (still GBIF-compatible but has no active support)
Setting up hosting and publishing organizations
In most cases, these protocols don’t generate Darwin Core Archives. Instead, the GBIF system uses the defined protocol to query the data page by page and parse the result. For example this is the result you get from querying the BoBO - Botanic Garden and Botanical Museum Berlin Observations (scientific name >= ‘Zaa’). This works pretty well for small datasets but can sometimes give inconsistent results for very large datasets as the ingestion process is much longer (days) and is more likely to be interrupted.
Although the BioCASe protocol does not generate Darwin Core Archives, the tool BioCASe Provider Software can be configured to expose ABCD-A (also called “BioCASe XML Archive”) and Darwin Core Archive and GBIF support both formats. In other words, data providers can choose how to share their data on GBIF (either with the BioCASe protocol or in an archive). Check out the BioCASe Provider Software documentation for more information.
In any case, the registration of these systems in GBIF is handled by the GBIF Secretariat.
This means that if you set up a BioCAse installation for example, you need to write to helpdesk@gbif.org and we will register the installation endpoint with the correct hosting organization. Afterwards, we will launch the “synchronization” of this installation. Depending on the way the installation and wrappers are configures, this will create the corresponding datasets on GBIF under the same organization. If the hosting organization differs from the publishing organization, you will have to notify helpdesk@gbif.org and we will make the update on our side manually.
To go further
Here is a little summary of all the tools I link in this post:
The chart below might give you an idea of how these tools are represented on GBIF. Note that some of them are still registered on GBIF but no longer host any datasets while some other host many datasets.
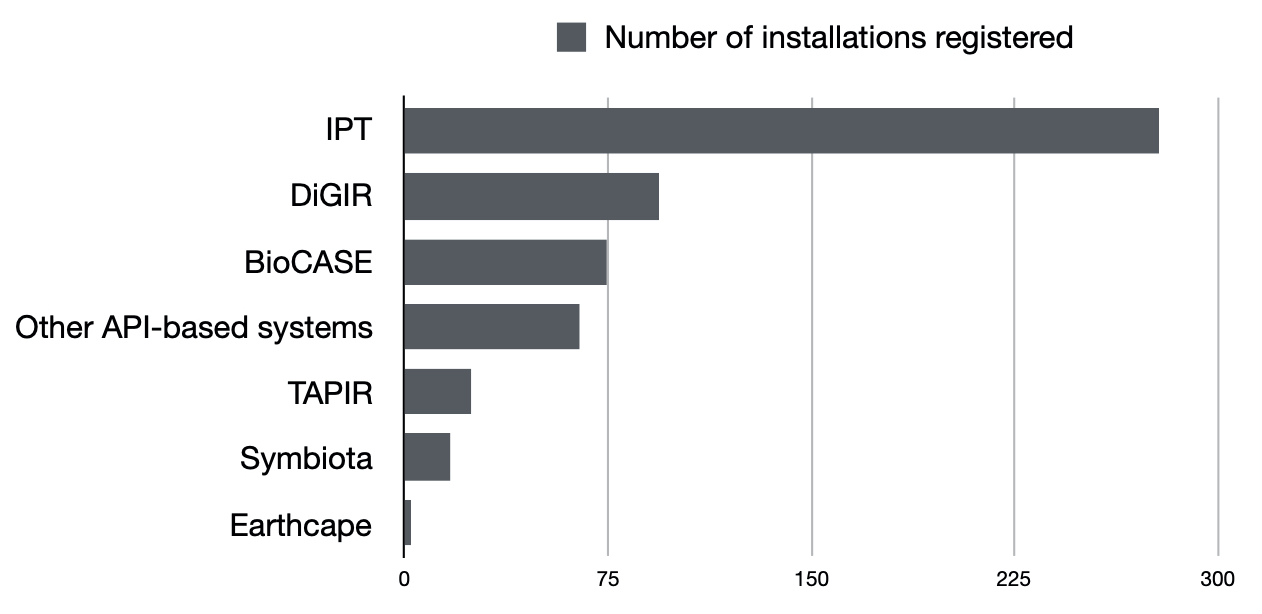
Please let me know if you know if I forgot to include any GBIF-compatible system in this post.
Should we make an other blogpost to compare these tools? If so, which criteria would you like to see represented?
